—
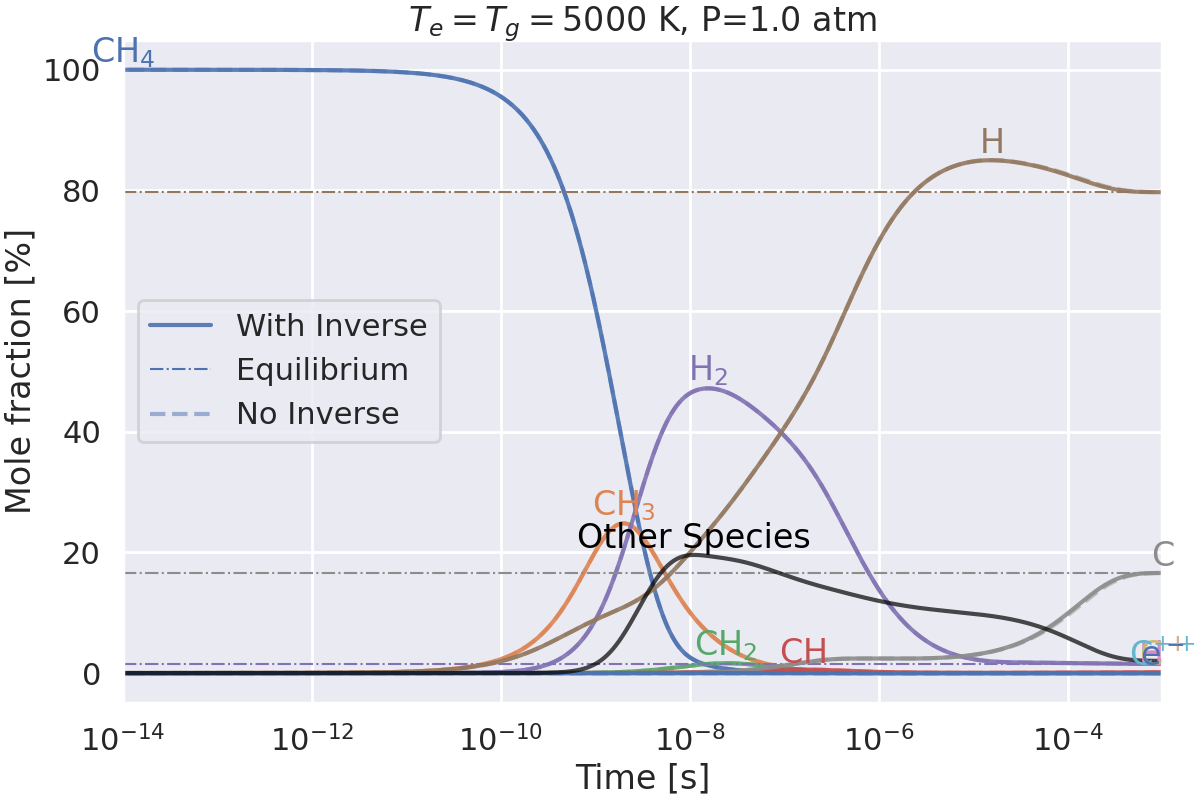
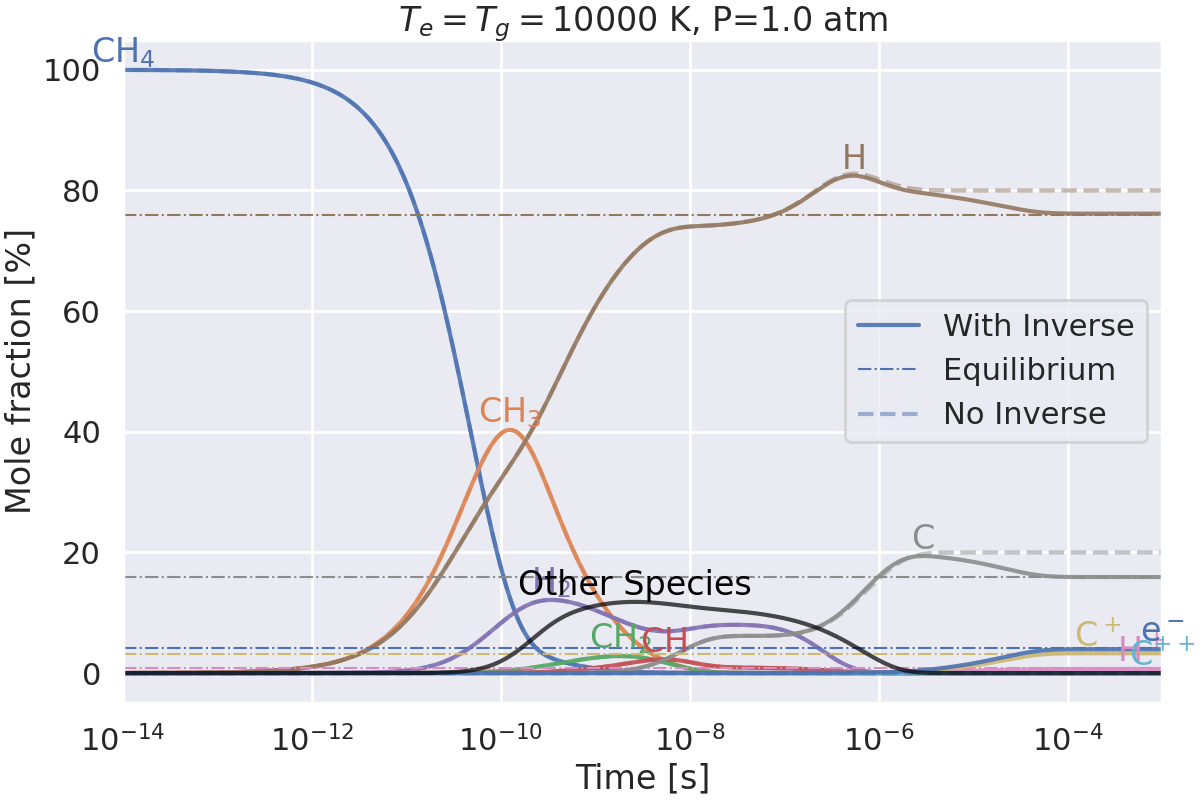
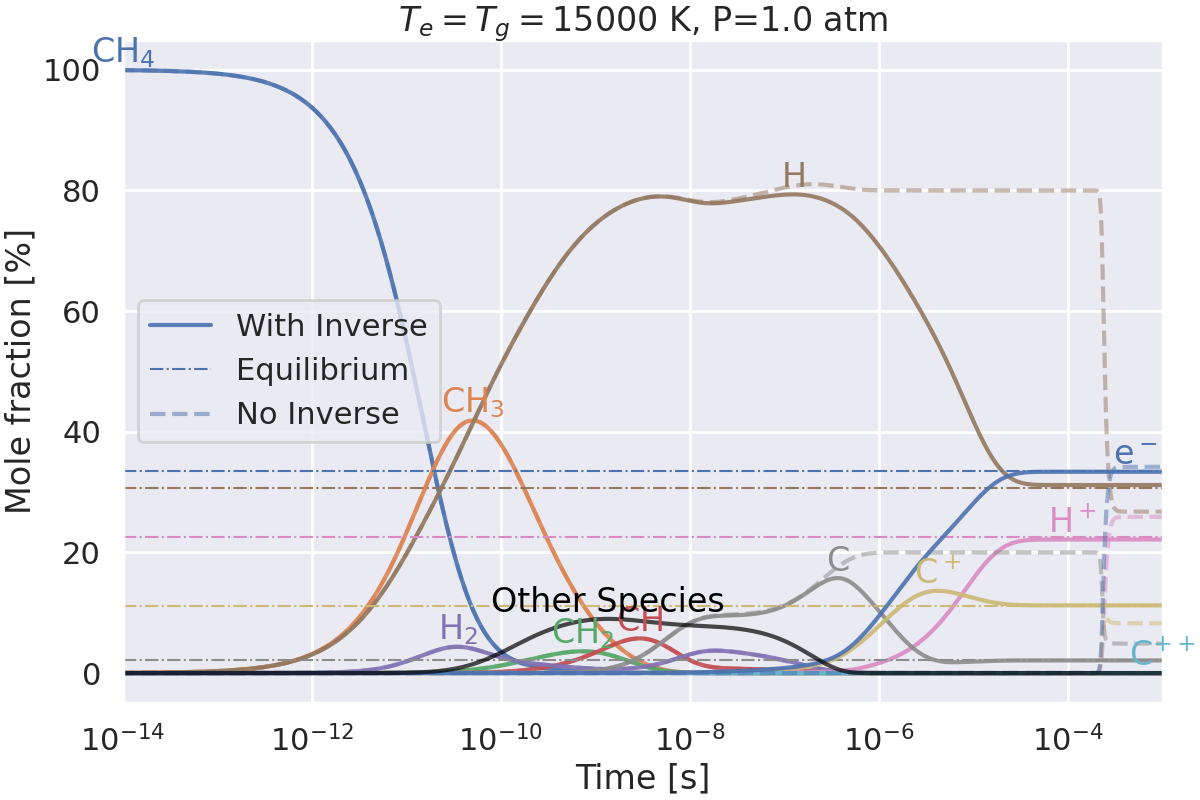
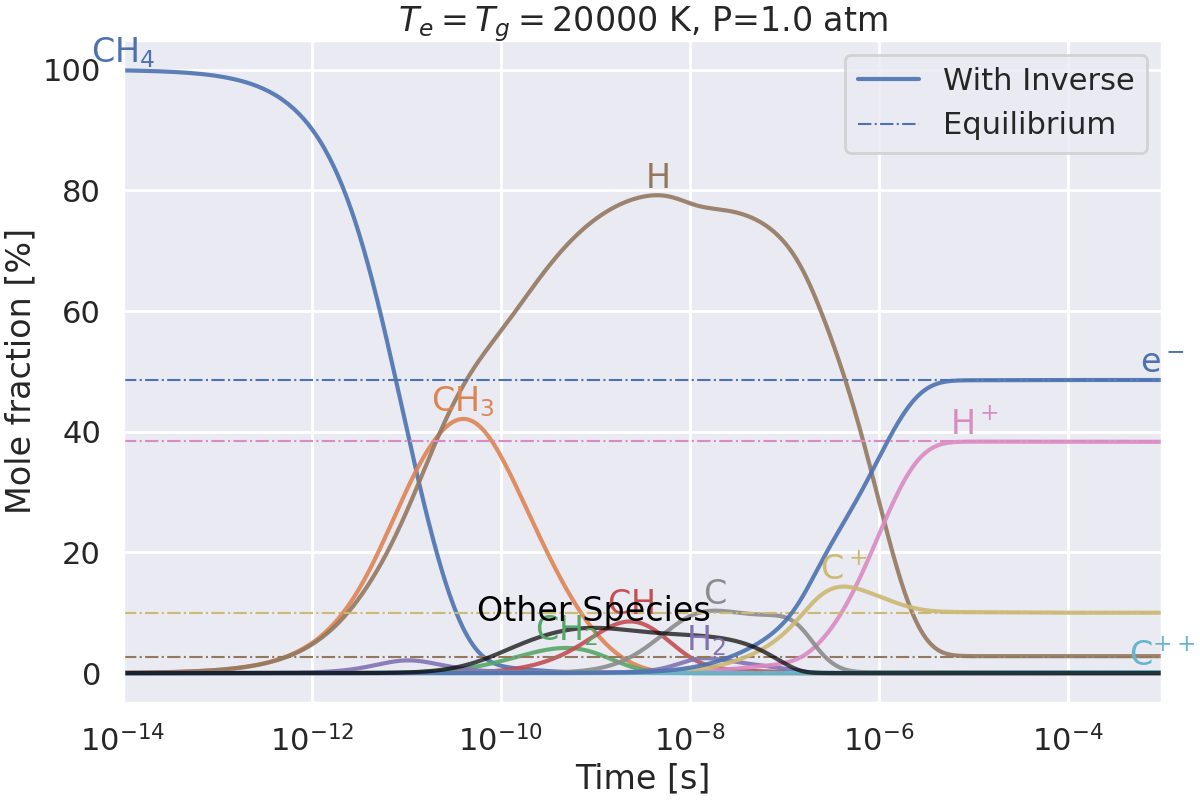
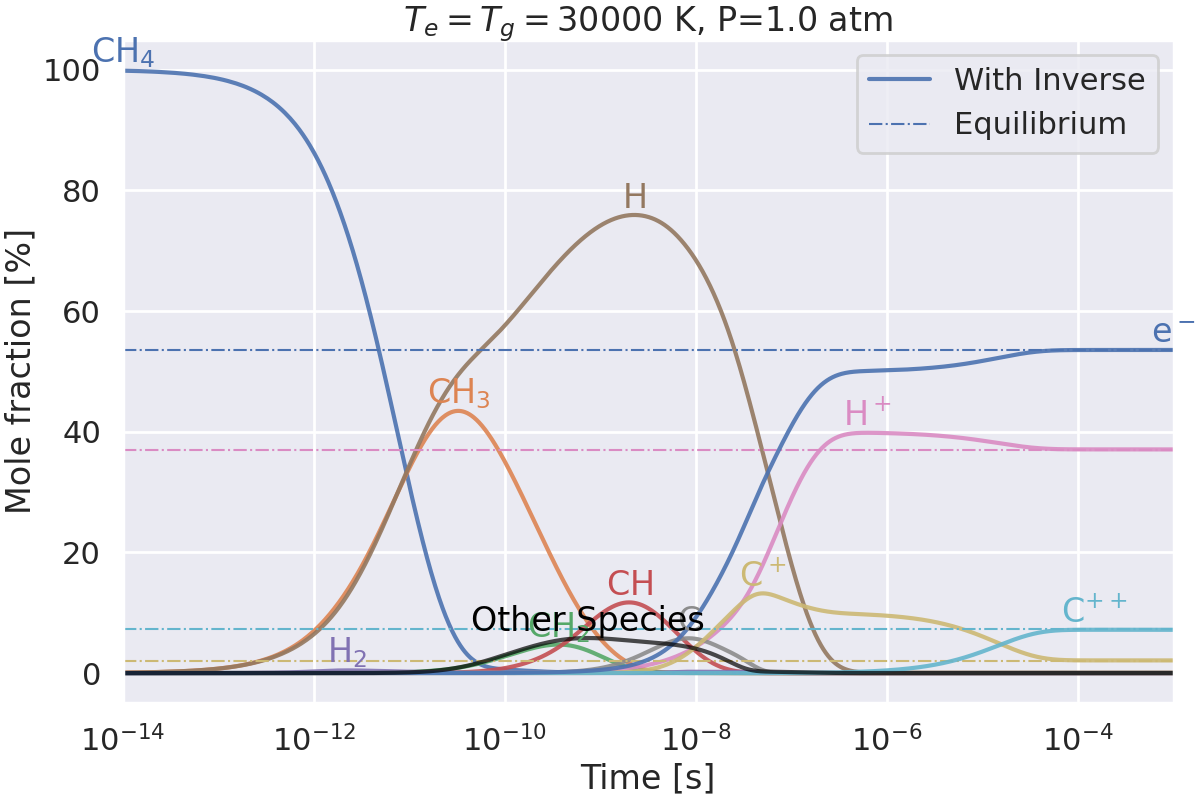
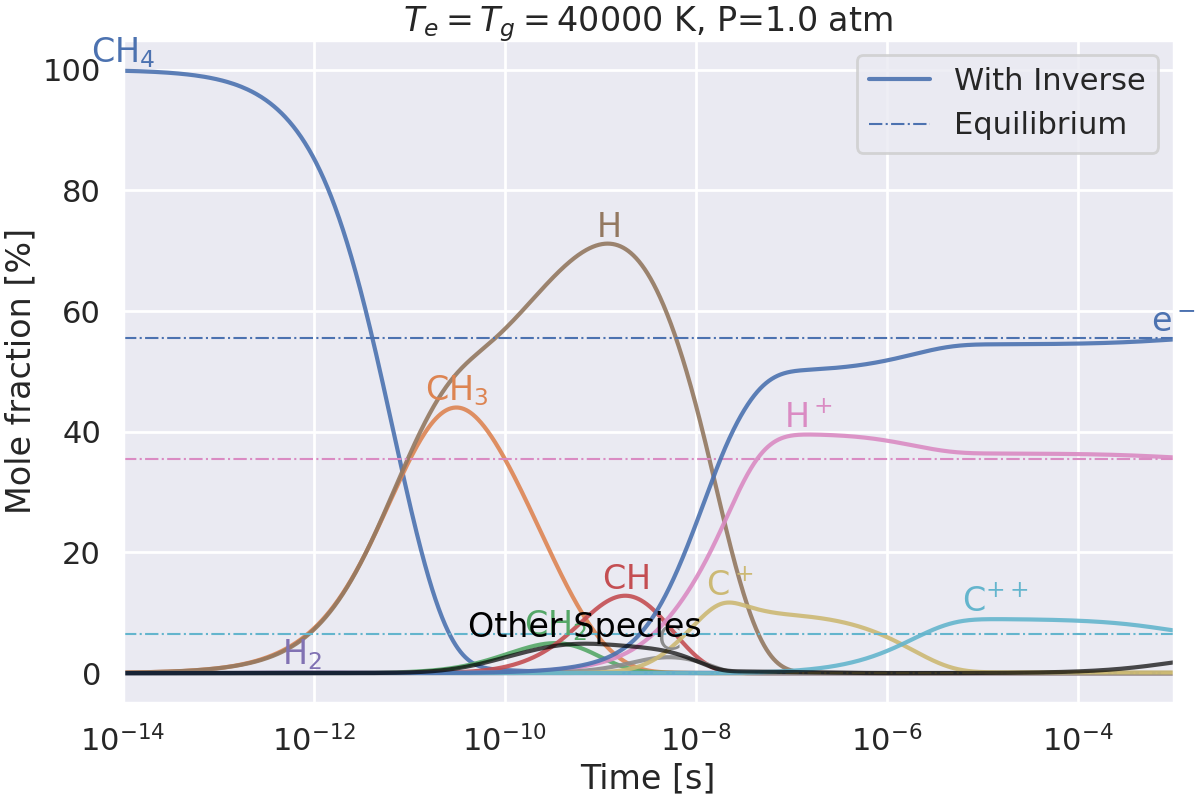
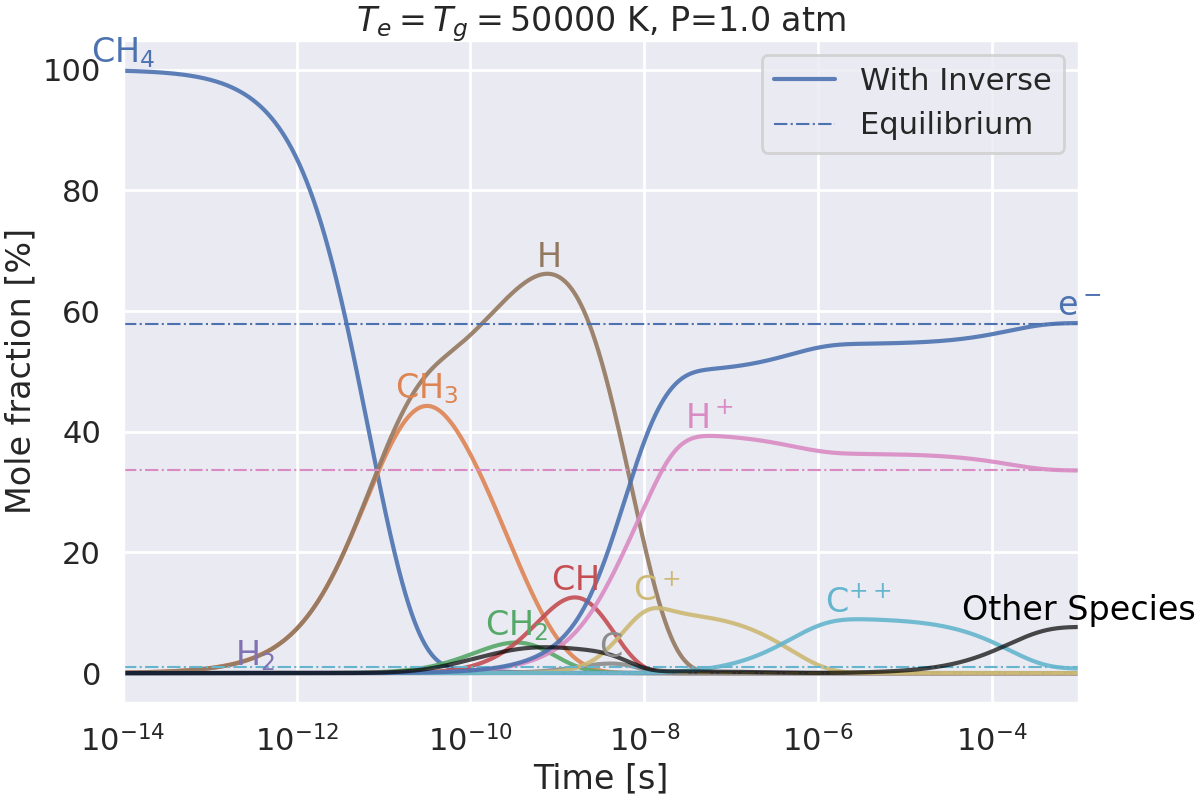
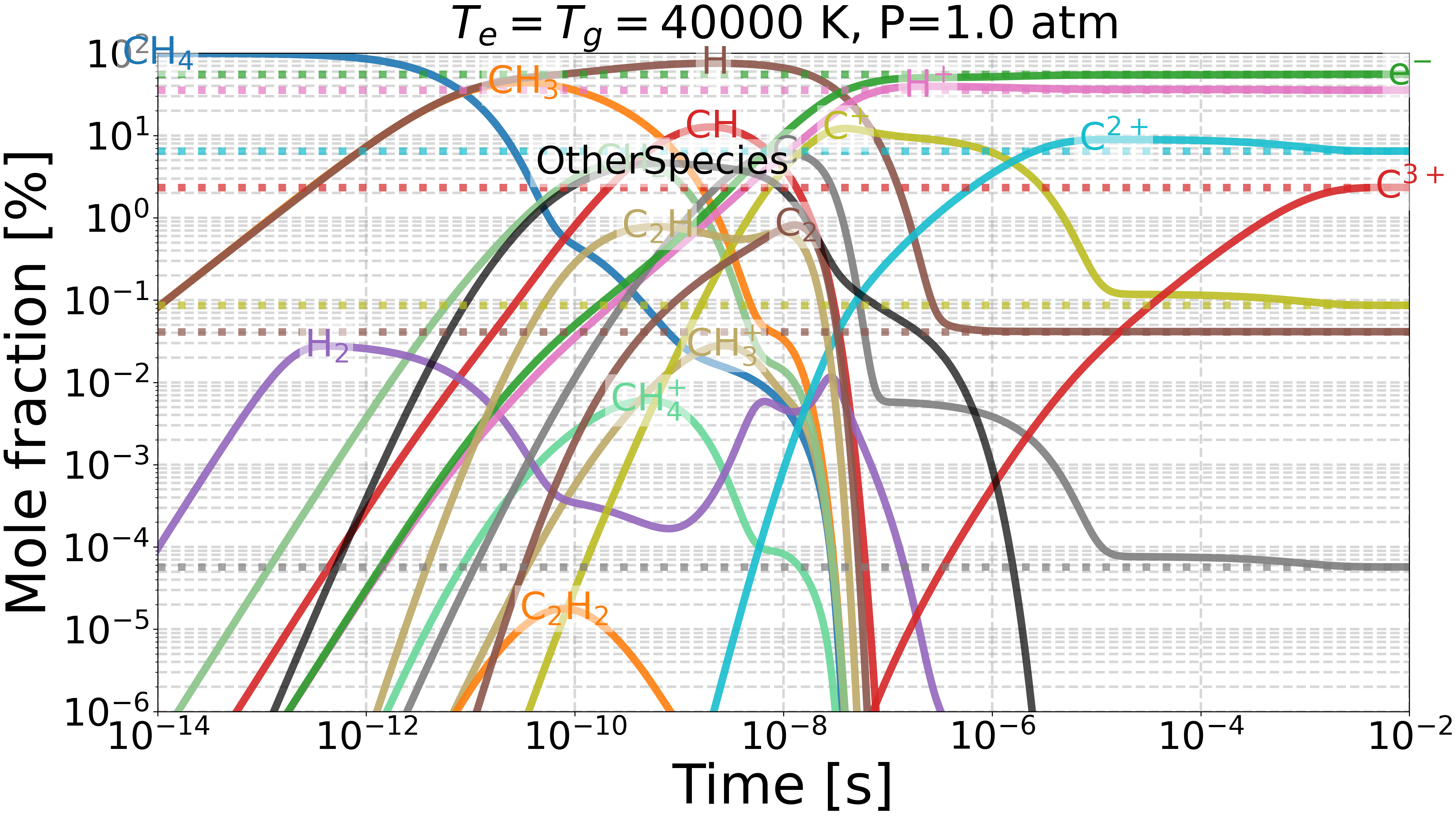
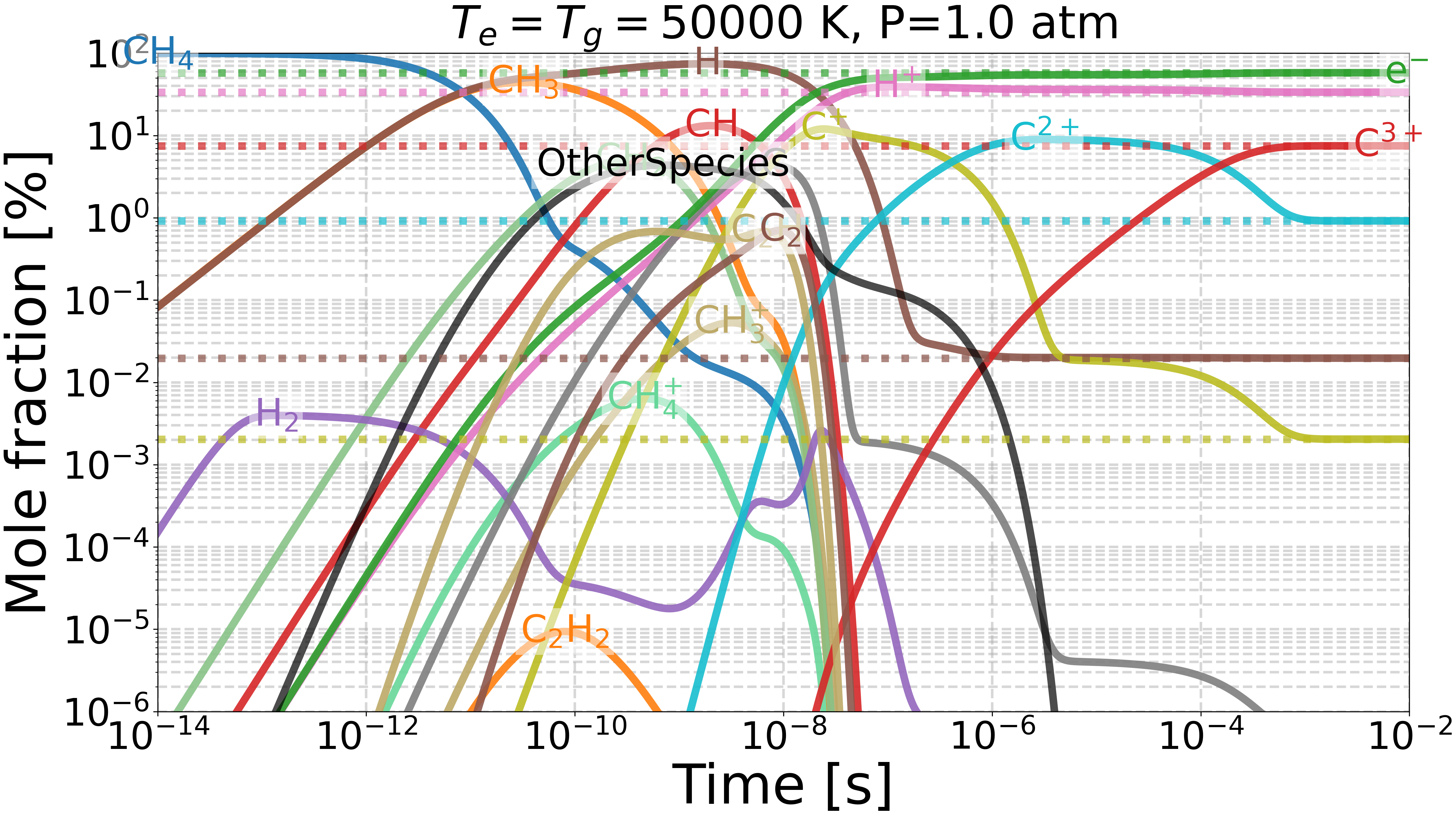
Do we get the same equilibrium with and without inverse reactions?#
This example checks if the equilibrium of two mechanisms is the same:
one with inverse reactions,
one without inverse reactions.
Key formula used for reverse rate constant calculation:
\[k_b = \frac{k_f}{K_{eq}}\]
where:
\(k_b\): backward rate constant
\(k_f\): forward rate constant
\(K_{eq}\): equilibrium constant
The equilibrium constant is computed from Gibbs free energy:
\[K_{eq} = \exp\left(-\frac{\Delta G}{R T}\right)\]
where:
\(\Delta G\): Gibbs free energy change
\(R\): universal gas constant
\(T\): temperature
For two-temperature plasma reactions, in equilibrium, the electron temperature is equal to the gas temperature, and the rate constant is computed using the above formula.
# This is an option for the online documentation, so that each image is displayed separately.
# sphinx_gallery_multi_image = "single"
Import the required libraries.#
from dataclasses import dataclass, field
import cantera as ct
import matplotlib.pyplot as plt
import numpy as np
from rizer.kin.extensible_rate import * # noqa: F403
from rizer.misc.plt_utils import get_species_in_latex, set_mpl_style
from rizer.misc.utils import get_path_to_data
set_mpl_style(nb_columns=1)
Define mechanisms and parameters.#
mechanism_no_inverse = get_path_to_data(
"mechanisms",
"Goutier2025",
"builder",
"CH4_to_C2H2_forward_reactions.yaml",
)
mechanism_with_inverse = get_path_to_data(
"mechanisms",
"Goutier2025",
"CH4_to_C2H2.yaml",
)
gas_no_inverse = ct.Solution(mechanism_no_inverse, name="gas")
gas_with_inverse = ct.Solution(mechanism_with_inverse, name="gas")
# gas_with_states = ct.Solution(
# mechanism_with_inverse, name="gas_with_electronic_excited_states"
# )
# Temperatures in Kelvin.
# For the mechanism with inverse reactions, we can go up to 50000 K,
# because the equilibrium is reached and the results are reliable.
temperatures_with_inverse = np.array(
[
1000,
2000,
5000,
10000,
15000,
20000,
30000,
40000,
50000,
]
)
# For the mechanism without inverse reactions, we cannot go higher than 15000 K,
# because the equilibrium is not reached and the results are not reliable.
temperatures_no_inverse = temperatures_with_inverse[temperatures_with_inverse < 15000]
# Pressure in Pascals.
P = ct.one_atm
# Initial mole fraction (here, we start with pure methane).
X_0 = "CH4:1"
# Species to plot, with their corresponding colors.
species_to_plot = {
"CH4": "#1f77b4",
"CH4+": "#67d799",
"CH3": "#ff7f0e",
"CH3+": "#bda967",
"CH2": "#89c389",
"CH": "#d62728",
"H2": "#9467bd",
"H": "#8c564b",
"H+": "#e377c2",
"C": "#7f7f7f",
"C(1D)": "#1f77b4",
"C+": "#bcbd22",
"C++": "#17becf",
"C+++": "#d62728",
"e-": "#2ca02c",
"C2H2": "#ff7f0e",
"C2H": "#bda967",
"C2": "#8c564b",
}
# Parameters for the reactor network.
simulation_time = 1e-2 # Simulation time in seconds
rtol = 1e-11 # Relative tolerance for the reactor network
max_steps = 100_000 # Maximum steps for the reactor network
max_time_step = 1e-5 # Maximum time step for the reactor network
Define the state groups for the different mechanisms.#
@dataclass
class StateGroup:
solution: ct.Solution
temperatures: np.ndarray
linestyle: str
linewidth: float
states: list[ct.SolutionArray] = field(default_factory=list)
no_inverse = StateGroup(
solution=gas_no_inverse,
temperatures=temperatures_no_inverse,
linestyle="--",
linewidth=5,
)
with_inverse = StateGroup(
solution=gas_with_inverse,
temperatures=temperatures_with_inverse,
linestyle="-",
linewidth=6,
)
# with_electronic_excited_states = StateGroup(
# solution=gas_with_states,
# temperatures=temperatures_with_inverse,
# linestyle="-.",
# linewidth=5,
# )
all_states: list[StateGroup] = [
no_inverse,
with_inverse,
# with_electronic_excited_states,
]
Compute equilibrium and time evolution for each mechanism and each temperature.#
# Compute the equilibrium for the mechanism with inverse reactions.
states_eq = ct.SolutionArray(gas_with_inverse, shape=temperatures_with_inverse.shape)
states_eq.TPX = temperatures_with_inverse, P, X_0
states_eq.equilibrate("TP")
# Compute the time evolution for each mechanism and each temperature.
for state_group in all_states:
print(f"Running equilibrium for {state_group.solution.name} mechanism")
for T in state_group.temperatures:
print(f"Running equilibrium for T={T} K")
# Initialize the reactor network for the current temperature.
state_group.solution.TPX = T, P, X_0
r1 = ct.IdealGasConstPressureReactor(state_group.solution, energy="off")
sim = ct.ReactorNet([r1])
sim.rtol = rtol # Set relative tolerance for the reactor network.
sim.max_time_step = max_time_step
sim.reinitialize()
# Store the states in a SolutionArray.
states = ct.SolutionArray(state_group.solution, 1, extra={"t": [0.0]})
i = 0
while sim.time < simulation_time:
# Advance the reactor network by one step.
sim.step()
# Append the current state to the SolutionArray.
states.append(state_group.solution.state, t=sim.time) # type: ignore
i += 1
if i % 1000 == 0:
print(f"{sim.time:.2e} / {simulation_time:.2e}")
state_group.states.append(states)
Running equilibrium for gas mechanism
Running equilibrium for T=1000 K
/home/runner/work/rizer/rizer/examples/kinetics/plot_equilibrium_reverse_reaction.py:198: DeprecationWarning: ReactorBase.__init__: After Cantera 3.2, the default value of the `clone` argument will be `True`, resulting in an independent copy of the `phase` being created for use by this reactor. Add the `clone=False` argument to retain the old behavior of sharing `Solution` objects.
r1 = ct.IdealGasConstPressureReactor(state_group.solution, energy="off")
9.89e-03 / 1.00e-02
Running equilibrium for T=2000 K
1.08e-04 / 1.00e-02
2.41e-03 / 1.00e-02
Running equilibrium for T=5000 K
5.89e-08 / 1.00e-02
2.57e-06 / 1.00e-02
1.56e-03 / 1.00e-02
Running equilibrium for T=10000 K
4.45e-09 / 1.00e-02
3.71e-07 / 1.00e-02
6.02e-06 / 1.00e-02
6.83e-03 / 1.00e-02
Running equilibrium for gas mechanism
Running equilibrium for T=1000 K
9.89e-03 / 1.00e-02
Running equilibrium for T=2000 K
1.00e-04 / 1.00e-02
2.50e-03 / 1.00e-02
Running equilibrium for T=5000 K
5.89e-08 / 1.00e-02
2.57e-06 / 1.00e-02
1.14e-03 / 1.00e-02
Running equilibrium for T=10000 K
4.45e-09 / 1.00e-02
3.34e-07 / 1.00e-02
5.23e-06 / 1.00e-02
1.17e-03 / 1.00e-02
Running equilibrium for T=15000 K
1.78e-09 / 1.00e-02
8.56e-08 / 1.00e-02
2.25e-06 / 1.00e-02
1.37e-03 / 1.00e-02
Running equilibrium for T=20000 K
1.01e-09 / 1.00e-02
4.30e-08 / 1.00e-02
1.04e-06 / 1.00e-02
2.24e-03 / 1.00e-02
Running equilibrium for T=30000 K
3.35e-10 / 1.00e-02
6.46e-09 / 1.00e-02
1.07e-07 / 1.00e-02
5.33e-05 / 1.00e-02
8.04e-03 / 1.00e-02
Running equilibrium for T=40000 K
1.33e-10 / 1.00e-02
2.84e-09 / 1.00e-02
2.75e-08 / 1.00e-02
2.82e-07 / 1.00e-02
2.44e-04 / 1.00e-02
Running equilibrium for T=50000 K
6.49e-11 / 1.00e-02
1.75e-09 / 1.00e-02
9.55e-09 / 1.00e-02
3.31e-08 / 1.00e-02
1.16e-06 / 1.00e-02
2.76e-03 / 1.00e-02
Plotting the results.#
for i, T in enumerate(temperatures_with_inverse):
print(f"Plotting results for T={T} K")
fig, ax = plt.subplots()
# Storage for the maximum mole fraction with the corresponding time of each species across all states.
max_fractions = {species: (0.0, 0.0) for species in species_to_plot}
max_fractions["Other Species"] = (0.0, 0.0)
for j, state_group in enumerate(all_states):
if i >= len(state_group.temperatures):
continue
states = state_group.states[i]
other_species = np.ones_like(states.t) * 100.0 # type: ignore
for species, color in species_to_plot.items():
if species not in state_group.solution.species_names:
continue
# Plot the time evolution of the mole fraction.
x = states.t # type: ignore
y = states(species).X * 100
other_species -= y[:, 0]
ax.plot(
x,
y,
linestyle=state_group.linestyle,
linewidth=state_group.linewidth,
alpha=0.9,
color=color,
)
# Add the species name at the maximum point, with the same color as the line.
max_x = x[y.argmax()]
if max_x < 1e-14:
max_x = 1e-14
max_y = y.max()
# Update the maximum fraction and corresponding time for the species.
if j != 0:
# For the mechanism without inverse reactions, we cannot trust the results,
# so we skip the update of the maximum fraction.
if max_y > max_fractions[species][0]:
max_fractions[species] = (max_y, max_x)
# Plot the remaining species as "Other Species"
ax.plot(
states.t, # type: ignore
other_species,
linestyle=state_group.linestyle,
linewidth=state_group.linewidth,
alpha=0.7,
color="black",
)
# Add the species name at the maximum point, with the same color as the line.
max_x = states.t[np.argmax(other_species)] # type: ignore
if max_x < 1e-14:
max_x = 1e-14
max_y = np.max(other_species)
# Update the maximum fraction and corresponding time for "Other Species".
if max_y > max_fractions["Other Species"][0]:
max_fractions["Other Species"] = (max_y, max_x)
# Plot the equilibrium values as horizontal lines.
for species, color in species_to_plot.items():
if species not in states_eq.species_names:
continue
y_eq = states_eq[i](species).X * 100
ax.hlines(
y=y_eq,
xmin=1e-14,
xmax=simulation_time,
linestyle=":",
color=color,
alpha=0.7,
linewidth=6,
)
# Add a text label for the species, at the maximum of all the states.
for species, (max_y, max_x) in max_fractions.items():
if max_y < 1e-6 and species != "Other Species":
continue # Skip species with a maximum mole fraction below 5%.
ax.text(
max_x,
max_y,
get_species_in_latex(species),
color=species_to_plot.get(species, "black"),
zorder=10,
horizontalalignment="center",
verticalalignment="center",
bbox=dict(facecolor="white", alpha=0.5, edgecolor="none"),
)
ax.set_title(f"$T_e=T_g={T}$ K, P={int(P / ct.one_atm):.1f} atm")
ax.set_xlabel("Time [s]")
ax.set_xlim(left=1e-14, right=simulation_time)
ax.set_xscale("log")
ax.set_ylabel("Mole fraction [%]")
ax.set_ylim(bottom=1e-6, top=100)
ax.set_yscale("log")
plt.show()
Plotting results for T=1000 K
Plotting results for T=2000 K
Plotting results for T=5000 K
Plotting results for T=10000 K
Plotting results for T=15000 K
Plotting results for T=20000 K
Plotting results for T=30000 K
Plotting results for T=40000 K
Plotting results for T=50000 K
Total running time of the script: (0 minutes 13.749 seconds)