—
Plot molar heat capacity of N₂, CH₄, H₂ and Ar vs. temperature.#
This example demonstrates how to compute and plot the molar heat capacity at constant pressure (\(c_p\)) of N₂, CH₄, H₂ and Ar as a function of temperature, from 300 to 20 000 K.
At each temperature the mixture is brought to chemical equilibrium at
constant T and P starting from one mole of the pure species. The
resulting \(c_p\) is Cantera’s frozen-composition heat capacity
cp_mole evaluated at the equilibrium composition,
expressed in J/mol/K.
The thermo data (NASA 9-polynomial) come from the GRC database.
Note
The temperature range of CH₄ is limited to 6 000 K by its NASA 9 polynomial fit, while N₂, H₂ and Ar are available up to 20 000 K.
See also
Plot specific heat vs. temperature for different species and thermo data. – plots the single-species \(c_p\) across several thermo databases for C/H species.
Plot thermodynamic properties of H₂ vs. temperatures. – plots the mass heat capacity of H₂ in LTE conditions.
Import the required libraries.#
import cantera as ct
import matplotlib.pyplot as plt
import numpy as np
from rizer.misc.plt_utils import get_species_in_latex, set_mpl_style
from rizer.misc.utils import get_path_to_data
set_mpl_style()
Load species from the GRC NASA 9 database.#
The file grc_nasa9_all.yaml contains NASA 9-polynomial thermo data for a
wide range of atomic and molecular species, covering temperatures up to
20 000 K for most species.
path_to_thermo = get_path_to_data("mechanisms/thermo", "grc_nasa9_all.yaml")
all_species: list[ct.Species] = ct.Species.list_from_file(str(path_to_thermo))
species_by_name: dict[str, ct.Species] = {s.name: s for s in all_species}
Compute the equilibrium molar cp for each species.#
For each species a single-species ideal-gas is created and equilibrated at
each temperature using equilibrate().
cp_mole gives the frozen-composition molar
heat capacity [J/kmol/K] of the mixture. Dividing by 1 000 converts to
J/mol/K.
T_min = 300.0 # K
T_max = 20_000.0 # K
n_points = 1000
species_names = ["N2", "CH4", "H2", "Ar"]
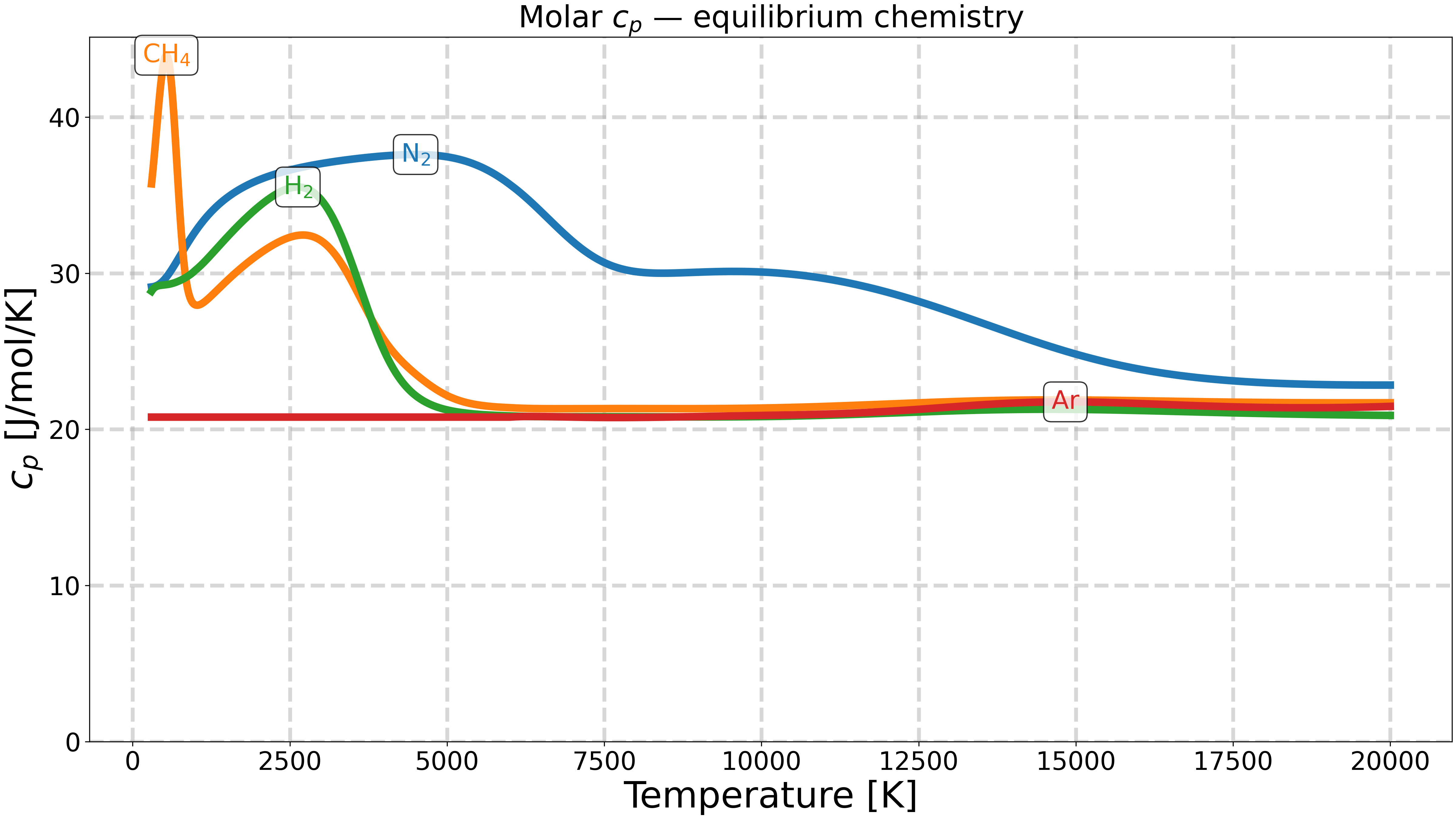
Equilibrium chemistry.#
The gas is equilibrated at each temperature before reading
cp_mole.
# To compute the equilibrium composition, we need to create a gas object that
# contains all the species. We load all the species from the GRC NASA 9
# database, then filter out any species that contain elements other than N, C, H
# and Ar to accelerate the computation.
all_species = ct.Species.list_from_file(str(path_to_thermo))
all_species = [
s for s in all_species if set(s.composition).issubset({"N", "C", "H", "Ar", "E"})
]
gas = ct.Solution(thermo="ideal-gas", kinetics="gas", species=all_species)
fig, ax = plt.subplots()
for name in species_names:
temperatures = np.linspace(T_min, T_max, n_points)
states = ct.SolutionArray(gas, shape=temperatures.shape)
states.TPX = temperatures, ct.one_atm, f"{name}:1"
states.equilibrate("TP")
cp_molar = states.cp_mole / 1e3 # J/kmol/K → J/mol/K
ax.plot(temperatures, cp_molar)
ax.text(
x=temperatures[np.argmax(cp_molar)],
y=np.max(cp_molar),
s=get_species_in_latex(name),
color=ax.get_lines()[-1].get_color(),
ha="center",
va="center",
bbox=dict(boxstyle="round", facecolor="white", alpha=0.8),
)
ax.set_xlabel("Temperature [K]")
ax.set_ylabel(r"$c_p$ [J/mol/K]")
ax.set_title(r"Molar $c_p$ — equilibrium chemistry")
ax.set_ylim(bottom=0)
plt.show()
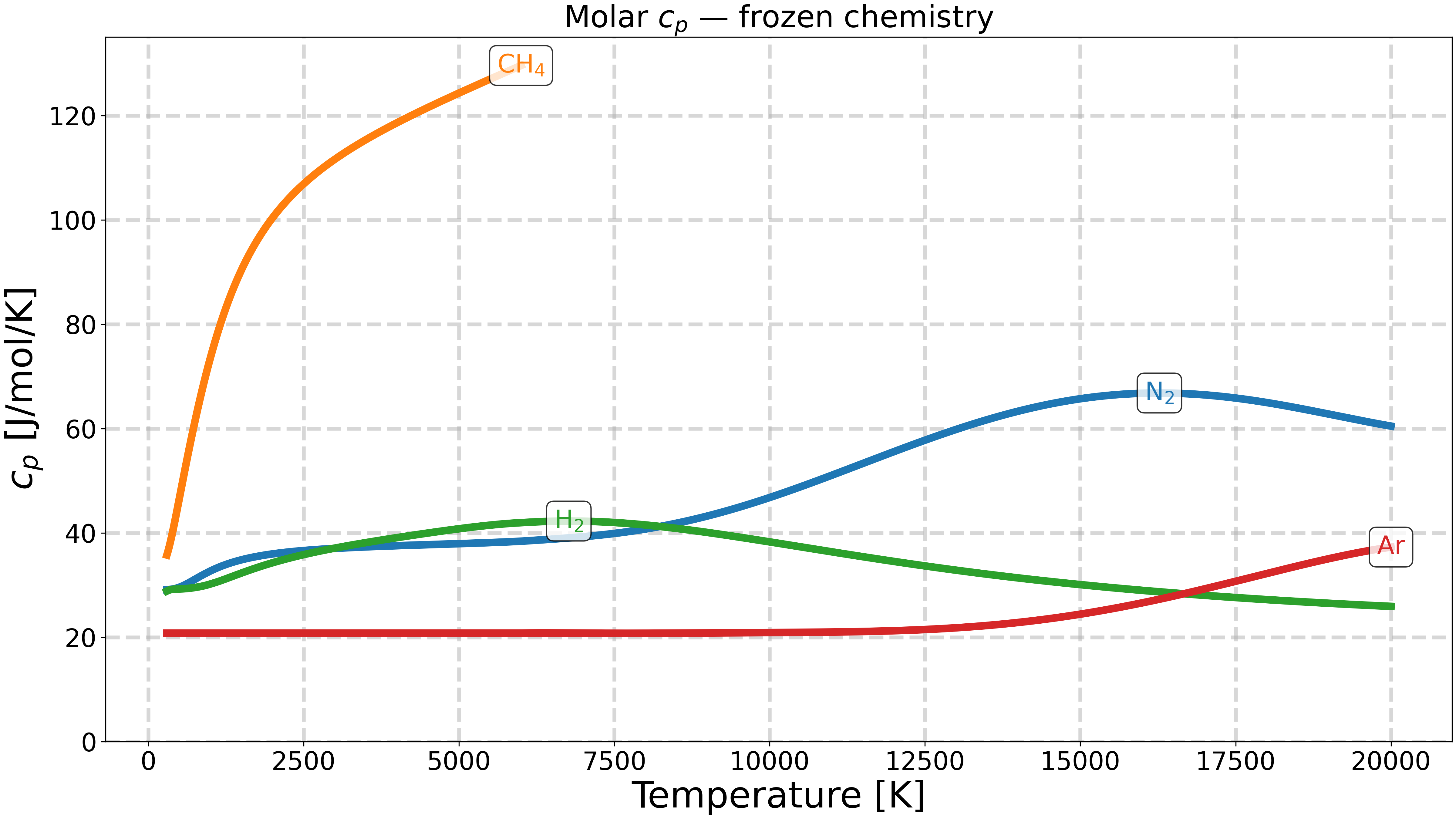
Frozen chemistry.#
The composition is kept fixed at the initial pure-species state (no
equilibration). cp_mole is then the pure
NASA-polynomial value for each species.
fig, ax = plt.subplots()
for name in species_names:
target_sp = species_by_name[name]
gas = ct.Solution(thermo="ideal-gas", kinetics="gas", species=[target_sp])
T_species_max = min(T_max, target_sp.thermo.max_temp)
temperatures = np.linspace(T_min, T_species_max, n_points)
states = ct.SolutionArray(gas, shape=temperatures.shape)
states.TPX = temperatures, ct.one_atm, f"{name}:1"
cp_molar = states.cp_mole / 1e3 # J/kmol/K → J/mol/K
ax.plot(temperatures, cp_molar)
ax.text(
x=temperatures[np.argmax(cp_molar)],
y=np.max(cp_molar),
s=get_species_in_latex(name),
color=ax.get_lines()[-1].get_color(),
ha="center",
va="center",
bbox=dict(boxstyle="round", facecolor="white", alpha=0.8),
)
ax.set_xlabel("Temperature [K]")
ax.set_ylabel(r"$c_p$ [J/mol/K]")
ax.set_title(r"Molar $c_p$ — frozen chemistry")
ax.set_ylim(bottom=0)
plt.show()
Total running time of the script: (0 minutes 14.979 seconds)
